- Blog
- Latest telugu christian songs youtube
- Android music making app
- Untangle firewall quota
- Microsoft flight simulator x com
- Stereology ihc axio
- Appserv php 5-3 1 download
- What is static ram
- What program do you use for live visuals
- Age of empires 2 pirates bay
- Nikon capture nx2 2-4-6 free full version
- Brazilian butt lift years later
- Places to sell osrs gold
- Empire earth 4 for mac
- Dubai arabic favo list xvi
- Mcmyadmin professional
- Laycock overdrive wiring diagram
- Autocad 2016 64 bit download with crack
- Microsoft project 2010 vs primavera p6
- Danganronpa killing harmony komaeda
- Fnaf 2 play online no download
- Amar chitra katha telugu free download
- Man of steel midi
- Foxit phantom help
- Pluraleyes 4 stuck premiere
- Football manager 2008 download
- Photoshop cc content aware fill
- Captain america the first avenger movie 2011

Several observations suggest that this neural specificity results from markedly lower levels of torsinB in neurons compared with non-neuronal cells. TorsinA null mice and mice homozygous for the DYT1 mutation exhibit neural-selective abnormalities of NE structure (NE ‘budding’) ( Goodchild et al., 2005 Kim et al., 2010). Their sequences are 68% identical and 85% similar, and they share cofactors LAP1 and LULL1 ( Ozelius et al., 1999 Goodchild and Dauer, 2005 Brown et al., 2014 Laudermilch et al., 2016). Prior work demonstrates conserved functions for torsinA and torsinB. The DYT1 mutation reduces protein stability and impairs interaction with cofactors (LAP1 and LULL1) that appear important for torsinA ATPase activity ( Goodchild and Dauer, 2005 Naismith et al., 2009 Zhao et al., 2013). Multiple lines of evidence demonstrate that the DYT1 mutation impairs torsinA function ( Tanabe et al., 2009 Liang et al., 2014 Weisheit and Dauer, 2015 Goodchild and Dauer, 2005). TorsinA is a nuclear envelope/endoplasmic reticulum (NE/ER) resident AAA+ protein ( ATPase Associated with diverse cellular Activities) ( Ozelius et al., 1997). These treatments suppress symptoms no therapies are based on disease pathogenesis or alter the emergence of symptoms. Treatments include deep brain stimulation, which is invasive, and anticholinergic drugs, which provide incomplete relief and are plagued by side effects ( Saunders-Pullman et al., 2002 Vidailhet et al., 2005).
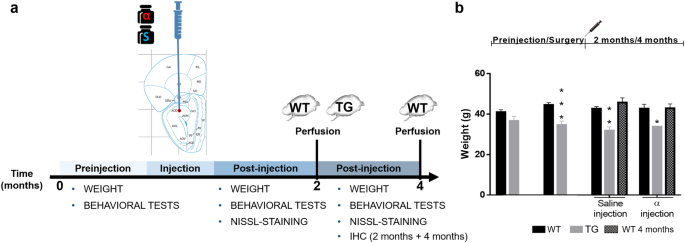
Only ~30% of mutation carriers exhibit symptoms, which vary in severity from mild to severely debilitating ( Albanese et al., 2011 Akbari et al., 2012). IntroductionĭYT1 dystonia is a dominantly inherited movement disorder that is caused by 3 bp in-frame deletion (ΔE mutation) in the TOR1A gene that encodes the torsinA protein ( Ozelius et al., 1997). These findings identify torsinB as a potent modifier of torsinA LOF phenotypes and suggest that augmentation of torsinB expression may retard or prevent symptom development in DYT1 dystonia. Reducing torsinB levels causes a dose-dependent worsening whereas torsinB overexpression rescues torsinA LOF-mediated abnormal movements and neurodegeneration. We demonstrate that torsinB levels bidirectionally regulate these phenotypes. Here, we examined whether torsinB expression levels impact the onset or severity of abnormal movements or neuropathological features in DYT1 mouse models. This work established that low neuronal levels of torsinB dictate the neuronal selective phenotype of nuclear membrane budding. Prior work demonstrates that torsinA and its paralog torsinB have conserved functions at the nuclear envelope. We explored this possibility in DYT1 dystonia, a neurodevelopmental movement disorder caused by a loss-of-function (LOF) mutation in the TOR1A gene encoding torsinA. We discuss the IHC methods necessary to utilize this analysis and provide detailed instruction on the use of QuPath in the pixel-based analysis of whole-slide images.Genetic redundancy can be exploited to identify therapeutic targets for inherited disorders. We compared swine brains that had undergone a closed head traumatic brain injury with brains of sham animals, and found a global increase in both microglial signal expression and phosphorylated tau. This method is fast, automated, unbiased, and easily replicable. The pixel-based method of analysis herein allows for statistical comparison of global protein expression between brains and generates heat-maps of stain intensity, visualizing stain signal across whole sections and permitting more specific investigation of regions of interest.
STEREOLOGY IHC AXIO FREE
Here we present the use of QuPath, a free and open source software, to quantify whole-brain sections stained with the immunohistochemical markers IBA1 and AT8, for microglia and phosphorylated tau respectively. Heterogeneous tissue types such as brain tissue have presented a further challenge to the development of automated analysis, as differing cellular morphologies result in either limited applicability or require large amounts of training tissue for machine-learning methods. Large scale unbiased quantification of immunohistochemistry (IHC) is time consuming, expensive, and/or limited in scope.
- Blog
- Latest telugu christian songs youtube
- Android music making app
- Untangle firewall quota
- Microsoft flight simulator x com
- Stereology ihc axio
- Appserv php 5-3 1 download
- What is static ram
- What program do you use for live visuals
- Age of empires 2 pirates bay
- Nikon capture nx2 2-4-6 free full version
- Brazilian butt lift years later
- Places to sell osrs gold
- Empire earth 4 for mac
- Dubai arabic favo list xvi
- Mcmyadmin professional
- Laycock overdrive wiring diagram
- Autocad 2016 64 bit download with crack
- Microsoft project 2010 vs primavera p6
- Danganronpa killing harmony komaeda
- Fnaf 2 play online no download
- Amar chitra katha telugu free download
- Man of steel midi
- Foxit phantom help
- Pluraleyes 4 stuck premiere
- Football manager 2008 download
- Photoshop cc content aware fill
- Captain america the first avenger movie 2011
